Morphology
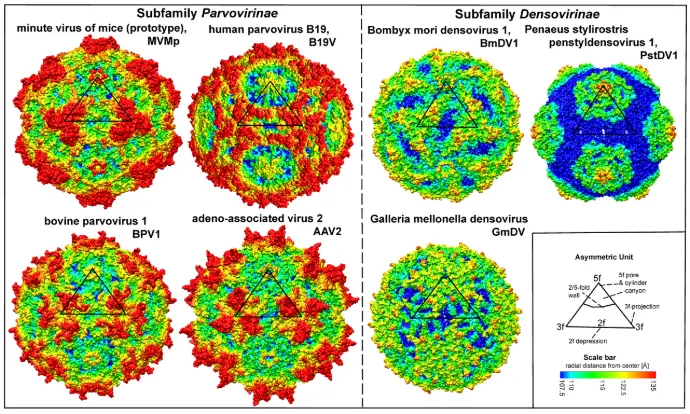
Virions are small, non-enveloped particles, 23–28nm in diameter, with no known essential lipids, carbohydrates, accessory proteins or histones. They are exceptionally rugged, often remaining infectious in the environment for months or years. X-ray crystallographic and cryoEM image reconstruction studies reveal a highly characteristic particle structure with T=1 icosahedral symmetry. Capsids are assembled from sixty structurally-equivalent polypeptide chains derived from the C-terminal end of a single capsid viral protein (VP) sequence. These monomers interdigitate extensively to create 60 asymmetric units per particle, which co-ordinate at twelve 5-fold, twenty 3-fold and thirty 2-fold axes (Figures 1.Parvoviridae and 2.Parvoviridae). Typical capsid surface features include depressions at each icosahedral two-fold axis, elevated protrusions surrounding each 3-fold axis, and raised cylindrical projections encircling each 5-fold vertex that are themselves surrounded by extensive canyon-like depressions. Such surface features are conspicuous in most family members, but their relative elevation varies between subfamilies and genera (Figure 1.Parvoviridae). Each 5-fold cylinder potentially encloses a central channel, which penetrates from the outer surface of the capsid to the particle interior. These serve as portals, mediating the entry and eventual exit of the viral genome from intact particles and in members of some genera, the phased extrusion of N-terminal regions from certain VPs that carry functional motifs required at specific stages in the viral life cycle. Some pathogenic shrimp and mosquito viruses (in genera Penstyldensovirus and Brevidensovirus respectively) have VP proteins that are relatively short and the loops connecting their beta-barrel strands appear minimal, resulting in particles with small diameters and a relatively smooth appearance (Figure:Parvoviridae). To date, structures of the VP1 N-terminal unique region have not been resolved at the atomic level, likely because their low copy number and/or flexibility makes them undetectable after rotational averaging in the context of the capsid. However, circular dichroism of adeno-associated virus 6 (AAV6) virus-like particles (VLPs) indicates that at neutral pH these regions adopt an α-helical conformation and unfold reversibly in a pH-dependent manner as the pH is lowered from 7.5 to 4.0. Around 20 nucleotides from the 5′-end of the viral genome may remain exposed at the surface of newly released virions, bearing a covalently-attached copy of the viral replication initiator protein, NS1, at its 5′-end. These are vestiges of the genome excision and packaging mechanisms (, and are vulnerable to enzymatic removal in the environment without affecting subsequent particle infectivity.
Virions are small, non-enveloped particles, 23–28nm in diameter, with no known essential lipids, carbohydrates, accessory proteins or histones. They are exceptionally rugged, often remaining infectious in the environment for months or years. X-ray crystallographic and cryoEM image reconstruction studies reveal a highly characteristic particle structure with T=1 icosahedral symmetry. Capsids are assembled from sixty structurally-equivalent polypeptide chains derived from the C-terminal end of a single capsid viral protein (VP) sequence. These monomers interdigitate extensively to create 60 asymmetric units per particle, which co-ordinate at twelve 5-fold, twenty 3-fold and thirty 2-fold axes (Figures 1.Parvoviridae and 2.Parvoviridae). Typical capsid surface features include depressions at each icosahedral two-fold axis, elevated protrusions surrounding each 3-fold axis, and raised cylindrical projections encircling each 5-fold vertex that are themselves surrounded by extensive canyon-like depressions. Such surface features are conspicuous in most family members, but their relative elevation varies between subfamilies and genera (Figure 1.Parvoviridae). Each 5-fold cylinder potentially encloses a central channel, which penetrates from the outer surface of the capsid to the particle interior. These serve as portals, mediating the entry and eventual exit of the viral genome from intact particles and in members of some genera, the phased extrusion of N-terminal regions from certain VPs that carry functional motifs required at specific stages in the viral life cycle. Some pathogenic shrimp and mosquito viruses (in genera Penstyldensovirus and Brevidensovirus respectively) have VP proteins that are relatively short and the loops connecting their beta-barrel strands appear minimal, resulting in particles with small diameters and a relatively smooth appearance (Figure:Parvoviridae). To date, structures of the VP1 N-terminal unique region have not been resolved at the atomic level, likely because their low copy number and/or flexibility makes them undetectable after rotational averaging in the context of the capsid. However, circular dichroism of adeno-associated virus 6 (AAV6) virus-like particles (VLPs) indicates that at neutral pH these regions adopt an α-helical conformation and unfold reversibly in a pH-dependent manner as the pH is lowered from 7.5 to 4.0. Around 20 nucleotides from the 5′-end of the viral genome may remain exposed at the surface of newly released virions, bearing a covalently-attached copy of the viral replication initiator protein, NS1, at its 5′-end. These are vestiges of the genome excision and packaging mechanisms (, and are vulnerable to enzymatic removal in the environment without affecting subsequent particle infectivity.
Figure 1.Parvoviridae. Surface images showing capsid topologies of viruses from selected genera in subfamilies Parvovirinae and Densovirinae. Radial distances from the particle centre are colour coded (blue to red), as indicated on scale bar. An asymmetric unit diagram is labelled to indicate icosahedral 2-, 3- and 5- fold axes and the 2/5-fold wall. The position of a single asymmetric unit is indicated on each model. Atomic co-ordinates were obtained as follows: minute virus of mice (prototype) (Protoparvovirus, PDB ID 1Z14), human parvovirus B19 (Erythroparvovirus, PDB ID 1S58), bovine parvovirus 1 (Bocaparvovirus, PDB 4QC8), adeno-associated virus 2 (Dependoparvovirus, PDB 1LP3), Bombyx mori densovirus 1 (Iteradensovirus, PDB ID 3POS) , Penaeus stylirostris penstyldensovirus 1 (Penstyldensovirus, PDB 3N7X) , Galleria mellonella densovirus (Ambidensovirus, PDB 1DNV). Figures were generated using the program Chimera.
Each VP chain forms a core beta-barrel structure of at least 8 strands (labelled βB–βI in Figure 2.Parvoviridae) arranged as two adjacent beta sheets (that spell CHEF and BIDG, see Figure 2E.Parvoviridae), with the BIDG sheet forming the inner surface of the capsid shell. Individual beta strands are linked by loops of variable length, sequence, and conformation, most of which project towards the outer surface of the capsid and give individual viruses their unique surface topology. The apices of these loops are the most varied and defined as variable regions (VRs). Although these monomer structures can differ widely between members of different genera (cf. Figure 2.Parvoviridae A–D), for viruses within a genus they are typically highly conserved. This is illustrated for members of genus Protoparvovirus in Figure 2E.Parvoviridae, where the Cα positions of the core secondary structure elements (β sheets A–I and helix A) of viruses from 4 different species are completely superimposed, and even the variable loops between these conserved structures follow very similar topologies (Figure 2E.Parvoviridae). Thus in general, parvovirus capsid structures are significantly more conserved than their protein sequence identities suggest: for example, the capsid amino acids of minute virus of mice (prototype, MVMp) and porcine parvovirus (PPV) share around 52% sequence identity, but 97% of their alpha carbons occupy similar positions (Figure 2E.Parvoviridae), and even between PPV and the insect virus Galleria mellonella densovirus (GmDV, genus Ambidensovirus; cf. Figure 2.Parvoviridae C and E), 40% of capsid residues align despite sharing only 9% sequence identity. Accordingly, capsid structures serve as a strong indicator of parvovirus phylogenetic identity.
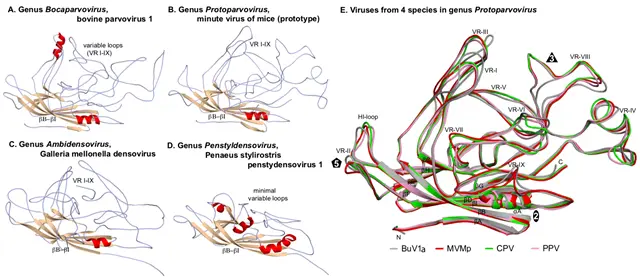
Figure 2.Parvoviridae. Folding of individual VPs in the parvovirus capsid shell. Capsid monomers from viruses in subfamilies Parvovirinae (A, B) and Densovirinae (C, D) are shown as a series of gray coils with β-sheets in brown and α-helices in red. β-barrels (βB–βI) and variable loops (VR I–IX) are indicated. (E) Superposed capsid protein monomers from viruses in 4 species of genus Protoparvovirus. Colour coded VP2 monomers from bufavirus 1a (human) (BuV1a; gray), minute virus of mice (prototype) (MVMp; red), canine parvovirus (CPV; green) and porcine parvovirus (PPV; pink) are overlaid, highlighting their conserved folds. The β-strands (βA–βI), VR regions (VR-I to VR-IX), and the HI-loop are shown and labeled. Icosahedral 2-, 3-, and 5-fold axes are represented by an oval, triangle, and pentagon, respectively. Atomic co-ordinates were as indicated in Figure 1.Parvoviridae, with the addition of PPV (PDB 1K3V), CPV (PDB 2CAS) and BuV1a (PDB 6BWX). Images were generated using PyMOL.
One consistent structural difference between virus capsids in members of the subfamilies Parvovirinae and Densovirinae involves the βA strand: whereas in vertebrate parvoviruses (Figure 2.Parvoviridae A–B and E) βA folds back and hydrogen bonds with βB from its own subunit, in densoviruses βB is a linear extension of the βA strand (Figure 2.Parvoviridae C–D), which allows βA to hydrogen bond with βB in the neighboring 2-fold related subunit, potentially enhancing the structural stability of this region. This is referred to as "domain-swapping".
Microbiology, Molecular Genetics & Immunology
Microbiology, Molecular Genetics, and Immunology are key areas of biological research that help us understand how living organisms function at the cellular and molecular levels. Microbiology studies microorganisms like bacteria, viruses, fungi, and protozoa, focusing on their structure, behavior, and role in the environment. Molecular Genetics looks at how genes work, how they are passed down, and how changes in genes can affect an organism. This includes studying DNA, RNA, and protein synthesis. Immunology examines how the immune system recognizes and responds to pathogens, while maintaining balance to avoid attacking the body’s own cells. These fields are essential for understanding biological processes and developing tools for research and industry.